|
Project
1: Retinal development and retinoblastoma |
|
Background:
The central nervous system (CNS)
consists of the brain, spinal cord and retina. Of these three
tissues, the retina is the easiest to work with because it is
well-characterized and relatively simple in structure. The retina
originates from precursor cells that have the potential of
differentiating into six classes of neuronal cells and one class of
glial cell. |
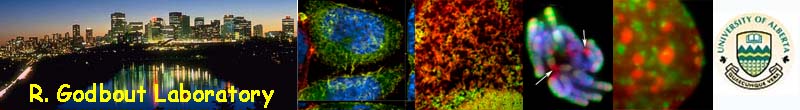
Dorsal expression of ALDH in
retina |
|
Goal:
To identify and characterize
genes involved in the differentiation of retinal precursor cells. We
are particularly interested in genes that function as transcriptional
regulators or as signaling molecules as they have the potential of
affecting the expression of numerous genes during retinal development
and in retinoblastoma. |
Experimental approach: We have used a variety of
screening procedures to identify genes that are differentially
expressed during retinal development and differentiation. One of the
genes that was identified using these screens is the transcription
factor AP-2. Based on expression analyses and in situ hybridizations,
AP-2 appears to be critical for the differentiation of retinal
precursor cells into two neuronal cell lineages called amacrine and
horizontal. We are using chromatin immunoprecipitation (ChIP) to
identify the target genes of AP-2. The role of AP-2 in retinoblastoma
is also under investigation. In a separate study, we have found that
the signaling molecule Disabled-1 (Dab1) exists in two forms in the
developing retina and brain: an early form restricted to precursor
cells and a late form restricted to differentiated cells. We are
studying the role of these two forms of Dab1 in the developing retina
and in retinoblastoma using mutagenesis assays, DNA transfection
experiments and expression analysis.

Retina tissue section
immunostained with AP-2 |
|
|
|
Project
2: Role of DEAD box proteins in retinoblastoma and retinal
development |
|
Background:
DEAD box proteins are putative RNA unwinding proteins that have been
implicated in all aspects of RNA metabolism (transcription, splicing,
processing, translation). We identified DEAD box 1 (DDX1) in a
differential screen of mRNAs expressed in retinoblastoma compared to
normal tissue. Subsequent experiments demonstrated that DDX1 was
amplified and over-expressed in a subset of retinoblastomas as well
as neuroblastomas, another type of childhood tumour.
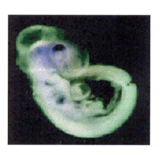
DEAD box proteins conserved
motifs
Goal:
To determine the function of DDX1 in retinoblastoma and in normal
retinal cells.
Experimental approach:
Using confocal microscopy and a biochemical approach, we have
demonstrated that DDX1 is primarily located in the nucleus where it
exists in close association with proteins implicated in RNA
transcription and processing. To further address the role of DDX1 in
normal and cancer cells, we are using a specialized
immunoprecipitation technique to identify the RNAs associated with
DDX1. We are also using the yeast two-hybrid system to identify
proteins that interact with DDX1. Transgenic mice carrying multiple
copies of the DDX1 gene have been generated and attempts are being
made to produce DDX1 gene knock-out mice. |
|
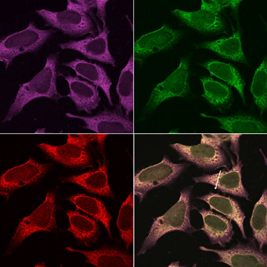
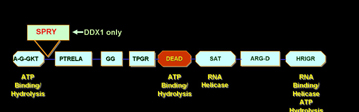
Immunofluorescence
analysis of DDX1 protein
|
|
|
Project 3:
Brain tumours and expression of glial cell
differentiation markers |
|
Background:
Malignant gliomas are brain tumours that are very
difficult to treat. Patients diagnosed with these
cancers usually die within two years of diagnosis. We
have found that a marker of glial cell differentiation
called brain fatty acid-binding protein (B-FABP) is
expressed in a subset of malignant glioma tumour cell
lines. Interestingly, B-FABP is coordinately expressed
with a second marker of glial cell differentiation,
called glial fibrillary acidic protein (GFAP).
|
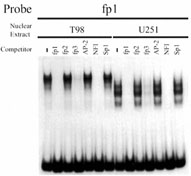
Gel shift assay |
|
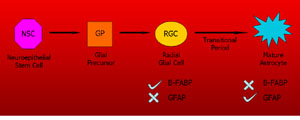
B-FABP and GFAP expression in
differentiating glial cells Goal:
To understand the
role that glial cell differentiation markers play in the
biology of malignant glioma tumours. |
Experimental approach:
We have identified malignant glioma cell lines that
express B-FABP and GFAP, and malignant glioma cell lines
that dont express these glial differentiation markers.
B-FABP has been introduced into the negative cell
lines, while an RNA interference approach was used to
reduce B-FABP levels in a positive cell line. We are
using these lines to address the role that B-FABP plays
in cellular growth properties such as proliferation
rate, growth in soft agar, invasiveness, motility, etc.
We are also using microchip cDNA arrays to find
differences between populations of cells that express B-FABP
compared to those that dont.
|
|
|