|
|
|
| Equipment | LSM710 | Multiphoton STED | Spinning Disc Confocal | DIMs | PALM | FCS | FLIM | TEM | Micro-Injection | HCS | Workstataions and Tools |
JEOL JEM-2100 Transmission Electron Microscope with GIF filter
The Facility is equipped with a 200kv JEOL 2100 Transmission Electron Microscope (TEM). TEM use an electron beam as light source (Lab6 crystal in this case) and the beam goes through (transmission) the specimen and the image is projected to an imaging device (e.g. fluorescence screen, film or a camera). The TEM can obtain resolution much higher than a light microscopy could offer (sub nm resolution versus ~200nm) due to the much shorter wavelength of the electron beam than visible light. Since electrons cannot penetrate thick specimen and electron light sources and beam requires vacuum in order to preventing arcing, TEM specimen must be specially processed before observing under TEM. For biological materials, this usually involves several steps:
- Fixation of the specimen which preserve the sample as close to the native status as possible before viewing under the TEM. This are usually achieved through one of the following methods: i) Chemical fixation which typically use cross linker molecules such as glutaraldehyde to fix major cellular components. ii) Cryofixation which involved in freezing the sample so rapidly that the samples are frozen in vitreous state. Some of the fixation reagent also enhance the contrast of the specimen under the TEM (e.g. OsO4)
- Contrast enhancing:since most biological material yield low contrast under electron beam, it is necessary to apply some heavy metal reagents to increase the contrast of the sample. Some of the commonly used one are OsO4 (for lipid staining), Uranyl acetate (for nucleus and negative staining); Lead citrate for general electron opaque contrast enhancing.
- Resin infiltration and plastic embedding: Since typical cellular structure is too thick to observe under TEM, it must be sectioned to thin sections (<100nm) prior viewing under TEM. This requires the specimen to be embedded in plastic resin. Typically, the specimen goes through steps like dehydration, resin infiltration and polymerization of resin (either in an oven or under UV light).
- Thin sectioning (ultra microtomy). Typical biological TEM has an acceleration voltage of 200kv or below and at this acceleration voltage, electrons can penetrate a specimen of thickness of less than ~200nm. A special instrument is used with a diamond knife to cut the specimen down to ~70-100nm for TEM observation and the sections are placed on to a specimen grid.
- Post staining if needed. Additional contrast enhancement is possible at this stage.
- Section stabilization:Many biological samples with resin are unstable under TEM beam (either due to local charging or heating generated by the electron beam). A thin layer of carbon (1-5nm) can be applied to the sections in order to increase the stability of the sections under the electron beam.
|
TEM images are formed by the interactions of electrons and the specimen and the TEM can operate in different modes to extract different information from the sample.
- Brightfield is the most common operation mode where most of the contrasts are formed by absorption of the electrons by the specimen yielding an image of different levels of shade which reflects the degree of electron absorptions of the sample.
- Darkfield mode explores the scattered electrons. A bright spot in the image means the structure is highly scattering (e.g. a gold particle).
- Electron Energy Loss Spectroscopy (EELS)detects the electrons which under inelastic interactions with the specimen (therefore loss some energy) and gates electrons of different energy level to the detector. Using this information, element composition of the specimen can be mapped. This mode is also very effective way to enhance contrast by filtering out the scattered electrons from the final image.
|
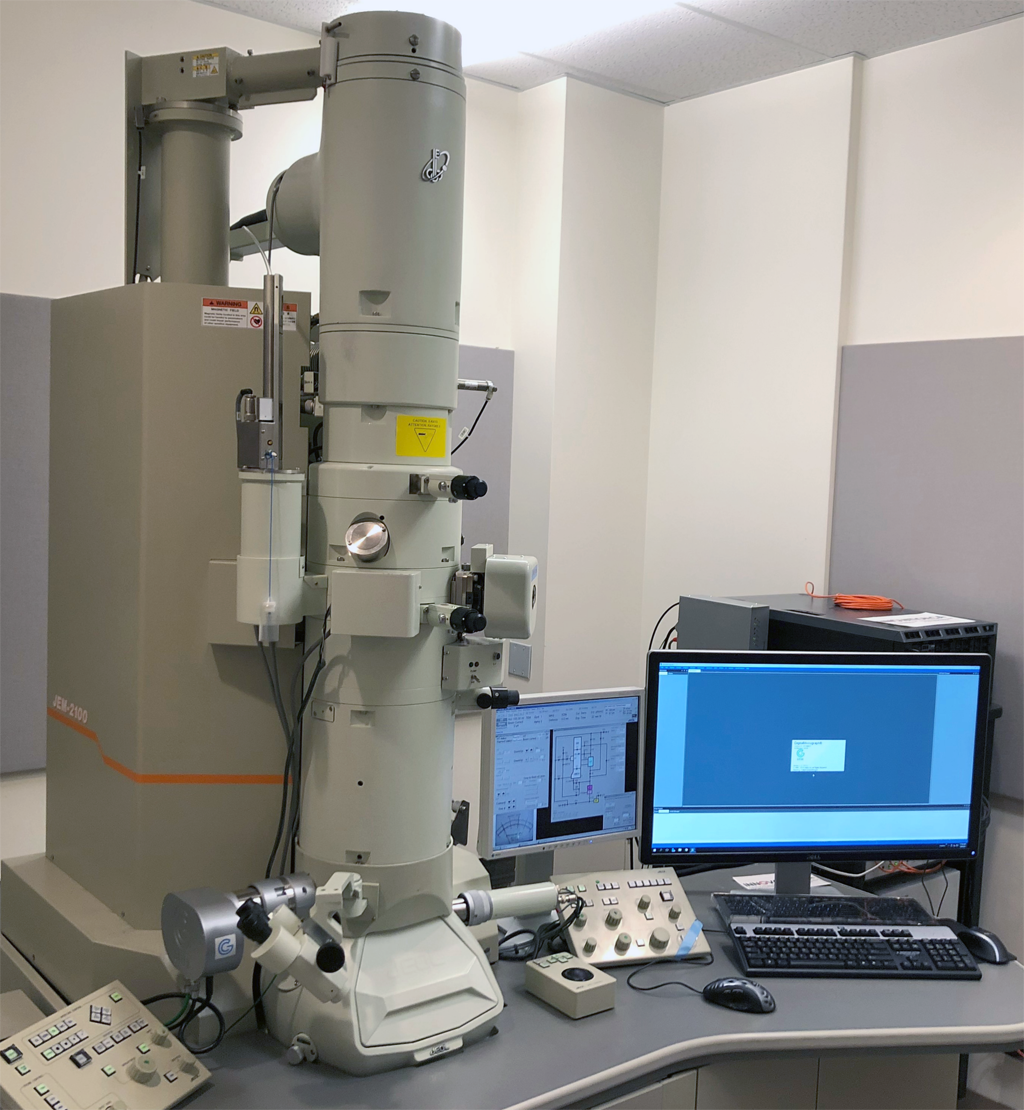 |
System Desciption:
The TEM in the Facility is updated in 2020 with a new GIF system and a direct electron detector. It is equipped with:
- A cryo sample holder for investigating sample at liquid nitrogen temperature.
- A double tilt sample holder for Tomography.
- A Gatan GIF Tridieum filter for EELS for element mapping.
- A high Angle Annular Dark field detector for dark field imaging.
- And some necessary TEM sample preparation for biological samples: These includes:
- High pressure freezer for cryofixation.
- Freeze substitution unit for substituting water from biological sample under cryo condition.
- Ultramicrotome with Cryo attachment for ultramicrotomy either at room temperature of low temperature.
- Immunogold labeler to perform immunogold labeling.
- Microwave automatic tissue processor for rapid tissue processing for TEM.
- Carbon coater with glowing discharge and metal shadowing capability.
- The scope is equipped with Gatan DigitalMicrograph software with Low-dose function for radiation sensitive sample (cryo specimen), Montaging for reconstruction of large field of view. For tomography, it uses SerialEM (http://bio3d.colorado.edu/SerialEM/)
|
TEM Applications.
TEM can be used to obtain details view of a specimen at atomic resolution level. For biological samples, TEM can give unambiguous identification of cellular compartments and organelles. In combination with specific molecular marker (e.g. antibody) and electron dense labels (immunogold particles or Nanoparticles), it is possible to identify association of a particular molecules with a specific organelle or with another type of molecules. With Tomography, 3-dimensional information can be obtained. With EELS, an elementary composition of the specimen can be mapped. For biological samples, for example, phosphorous rich and nitrogen rich domains can be mapped. The TEM is also equipped with cryo sample holder to observe sample under cryo-conditions.
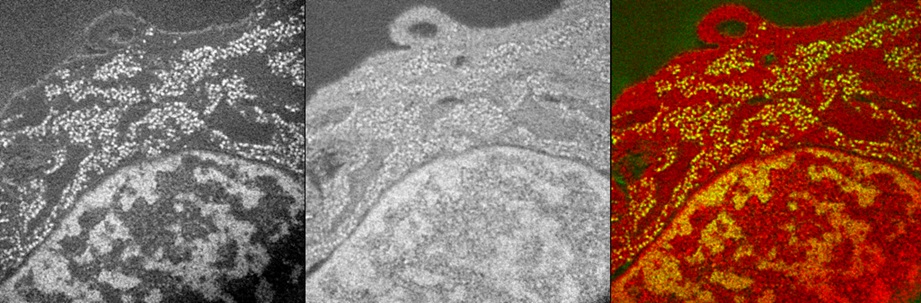
Electron Spectroscopic Imaging mouse cell nucleus (CH310T1/2) showing the Phosphorous map (backbone of nucleic acids (DNA & RNA) , Left) and Nitrogen map (middle) and the overlay of the 2 elements (color picture on the right) demonstrating the anatomic nuclear structures like the chromatin and the interchromatin compartment of the nucleus.
Images are provided by Dr. Hilmar Strickfaden, from Dr. Michael Hendzel lab.
|
. |
|
